小编use*_*211的帖子
使用BIC标准运行逐步线性模型
是否可以设置逐步线性模型以使用BIC标准而不是AIC?
我一直在尝试这个,但它仍然使用AIC值而不是BIC计算每一步
null = lm(data[,1] ~ 1)
full = lm(data[,1] ~ age + bmi + gender + group)
step(null, scope = list(lower=null,upper=full),
direction="both", criterion = "BIC")
推荐指数
解决办法
查看次数
使用电子表格数据在R中运行线性模型
我有一个由106个两种类型的人组成的数据集 - a和b有各种变量,例如年龄和性别.我想运行一个线性模型,根据协变量预测每个人是a型还是b型.
我使用以下方法读取每个人的年龄,性别和类型标签的值:
`data = read.xlsx("spreadsheet.xlsx",2, as.is = TRUE)`
age = data$age
gender = data$gender
type = data$type
每个都是以下形式:
age = [28, 30, 19, 23 etc]
gender = [male, male, female, male etc]
type = [a b b b]
然后我尝试使用以下方法设置模型:
model1 = lm(type ~ age + gender)
但我收到此错误消息:
Warning messages:
1: In model.response(mf, "numeric") :
using type="numeric" with a factor response will be ignored
2: In Ops.factor(y, z$residuals) : - not meaningful for factors
我尝试使用以下方法更改类型,年龄和性别的格式:
age = as.numeric(as.character(age))
gender …推荐指数
解决办法
查看次数
在qq图中添加置信区间?
有没有办法在qqplot中添加置信区间?
我有一个基因表达值的数据集,我用PCA可视化:
pca1 = prcomp(data, scale. = TRUE)
我现在通过检查数据与正态分布的分布来寻找异常值:
qqnorm(pca1$x,pch = 20, col = c(rep("red", 73), rep("blue", 33)))
qqline(pca1$x)
这是我的数据:
数据= [2.48 104 4.25 219 0.682 0.302 1.09 0.586 90.7 344 13.8 1.17 305 2.8 79.7 3.18 109 0.932 562 0.958 1.87 0.59 114 391 13.5 1.41 208 2.37 166 3.42]
我现在想绘制95%置信区间来检查哪些数据点在外面.关于如何做到这一点的任何提示?
推荐指数
解决办法
查看次数
将不同长度的向量保存在矩阵/数据帧中
我有一个称为长度为166860的数字区域。它由412个不同的元素组成,大部分长度为405,有些长度为809。我有它们的开始和结束ID。
我的目标是提取它们并将其放入具有412列的矩阵/数据框中
现在,我正在尝试以下代码:
m = matrix(NA,ncol=412, nrow=809)
for (j in 1:412){
temp.start = start.ids[j]
temp.end = end.ids[j]
m[,j] = area[temp.start:temp.end]
}
但是我最终得到以下错误消息:
“ m [,j] =区域[temp.start:temp.end]中的错误:要替换的项目数不是替换长度的倍数”
推荐指数
解决办法
查看次数
使用ComBat时出错
我是R的新手,我正在尝试在R sva库中使用331 x 89基因表达值矩阵的ComBat脚本.我的数据由5个批次组成,它以这种方式排序,因此前106行对应批次1,后者106对应批次2,依此类推.
batch1 <- rep(1,times=106)
batch2 <- rep(2,times=106)
batch3 <- rep(3,times=39)
batch4 <- rep(4,times=26)
batch5 <- rep(5,times=54)
batch.type <- as.factor(c(batch1,batch2,batch3,batch4,batch5))
然后我尝试使用此命令使用ComBat:
ComBat(data,batch=batch.type,mod=NULL)
我收到以下读数和错误消息:
"Found 5 batches
Found 0 categorical covariate(s)
Standardizing Data across genes
Error in solve(t(design) %*% design) %*% t(design) %*% t(as.matrix(dat)) :
non-conformable arguments"
推荐指数
解决办法
查看次数
nlme重复措施出错
我试图在57个不同的时间点运行一个重复测量的线性混合模型.但我不断收到错误消息:
Error in solve.default(estimates[dimE[1L] - (p:1), dimE[2L] - (p:1), drop = FALSE]) :
系统在计算上是单数的:倒数条件数= 7.7782e-18
这是什么意思?
我的代码是这样的:
model.dataset = data.frame(TimepointM=timepoint,SubjectM=sample,GeneM=gene)
library("nlme")
model = lme(score ~ TimepointM + GeneM,data=model.dataset,random = ~1|SubjectM)
这是数据:
score = c(2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,6,7,2,-3,11,14,1,7,6,7,2,-3,11,14,1,7,6,2,-3,11,14,1,7,7,2,-3,11,14,1,7,6,2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,6,2,-3,11,14,1,7,7,2,-3,11,14,1,7,6,7,2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,6,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,6,7,2,-3,11,14,1,7,6,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7,2,-3,11,14,1,7)
timepoint = c(1,1,1,1,1,1,2,2,2,2,2,2,3,3,3,3,3,3,3,4,4,4,4,4,4,5,5,5,5,5,5,6,6,6,6,6,6,7,7,7,7,7,7,8,8,8,8,8,8,8,9,9,9,9,9,9,10,10,10,10,10,10,10,12,12,12,12,12,12,12,12,13,13,13,13,13,13,13,13,14,14,14,14,14,14,14,14,15,15,15,15,15,15,16,16,16,16,16,16,16,16,17,17,17,17,17,17,18,18,18,18,18,18,19,19,19,19,19,19,19,20,20,20,20,20,20,21,21,21,21,21,21,24,24,24,24,24,24,24,25,25,25,25,25,25,25,27,27,27,27,27,27,28,28,28,28,28,28,29,29,29,29,29,29,30,30,30,30,30,30,30,31,31,31,31,31,31,31,32,32,32,32,32,32,33,33,33,33,33,33,33,33,34,34,34,34,34,34,34,35,35,35,35,35,35,36,36,36,36,36,36,36,37,37,37,37,37,37,38,38,38,38,38,38,39,39,39,39,39,39,39,40,40,40,40,40,40,40,41,41,41,41,41,41,41,41,42,42,42,42,42,42,42,42,43,43,43,43,43,43,44,44,44,44,44,44,44,45,45,45,45,45,45,46,46,46,46,46,46,47,47,47,47,47,47,48,48,48,48,48,48,49,49,49,49,49,49,49,50,50,50,50,50,50,51,51,51,51,51,51,52,52,52,52,52,52,52,53,53,53,53,53,53,53,54,54,54,54,54,54,55,55,55,55,55,55,56,56,56,56,56,56,57,57,57,57,57,57)
sample = c("S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S13T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S01T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0","S02T0","S03T0","S07T0","S09T0","S10T0","S12T0")
gene =c(24.1215870,-18.8771658,-27.3747309,-41.5740199,26.1561877,-2.7836332,20.8322796,36.5745088,-24.1541743,-11.2362216,4.9042852,7.4230219,155.8663563,16.4465366,-11.7982286,-1.6102783,-35.9559091,27.7909495,-13.9181661,-29.6037658,-68.4297261,-45.0877920,-48.3157529,17.1649982,-26.9084544,19.7358439,-5.8991143,-24.1541743,-23.5960654,13.0780939,-2.7836332,18.6394081,-28.3157487,-49.9186269,-33.7086648,41.6864242,-30.6199654,36.1823804,-36.5745088,-49.9186269,-44.9448864,-4.9042852,-34.3314764,62.3465425,-42.7609951,-11.7982286,-32.2055657,-56.1811080,5.7216661,-17.6296771,4.3857431,-43.6534459,9.6616697,-44.9448864,18.7997599,-12.9902884,109.1064494,7.6750504,-43.6534459,-17.7130611,-25.8433097,5.7216661,-18.5575548,35.2750175,36.1823804,2.3596457,-25.7644526,-55.0574858,15.5302365,-19.4854325,73.3687689,63.1668918,20.8322796,16.5175201,-22.5438960,-28.0905540,15.5302365,7.4230219,39.5062602,107.4657509,36.1823804,-23.5964573,-45.0877920,-43.8212642,4.0869043,-40.8266205,26.3375068,13.1572292,-25.9561030,-40.2569571,-52.8102415,2.4521426,-49.1775202,246.1047731,36.1823804,11.7982286,-35.4261223,-26.9669318,-2.4521426,-38.0429873,38.5656349,9.8679219,16.5175201,8.0513914,-42.6976421,26.9735686,-26.9084544,4.3857431,12.9780515,-32.2055657,-33.7086648,9.8085704,-36.2800196,215.7518511,6.5786146,-9.4385829,-19.3233394,-40.4503978,17.1649982,-7.4230219,14.2536650,-23.5964573,-53.1391834,-52.8102415,22.0692834,-54.7447866,24.1215870,-44.8332688,-24.1541743,-42.6976421,26.9735686,-40.8266205,191.1413737,17.5429723,-70.7893718,-37.0364006,-39.3267756,-4.9042852,-0.9278777,93.5198138,-6.5786146,-24.7762801,-28.9850091,-39.3267756,22.0692834,-50.1053979,14.2536650,23.5964573,-20.9336177,-53.9338637,14.7128556,-39.8987428,4.3857431,-64.8902575,-59.5802966,-33.7086648,22.0692834,2.7836332,46.0503024,-35.3946859,-43.4775137,-53.9338637,30.2430921,-34.3314764,80.3942259,28.5073300,-87.3068919,-24.1541743,-62.9228410,13.0780939,-25.0526990,35.0859447,-24.7762801,-38.6466789,-58.4283523,31.0604729,0.0000000,24.4562563,1.0964358,-27.1359259,-75.6830794,-16.8543324,20.4345217,-11.1345329,74.1390629,18.2282447,-27.3044720,-45.2890768,-46.7707724,15.3258912,-27.9523169,-6.9763039,117.3099418,18.6394081,-21.2368115,-38.6466789,-34.8322870,22.0692834,-48.2496425,6.5786146,-64.8902575,-51.5289052,-80.9007955,23.7040451,-26.9084544,223.1349942,8.7714862,10.6184058,-127.2119846,-31.4614205,0.8173809,-16.7017993,9.8679219,-35.3946859,-54.7494617,-44.9448864,14.7128556,-18.5575548,97.5827836,-166.3550237,-95.0064189,-123.5984376,104.6247509,-121.5519839,33.9895089,-44.8332688,-40.2569571,-56.1811080,51.4949946,0.0000000,-16.9312544,95.9808615,6.5786146,-21.2368115,-9.6616697,-13.4834659,10.6259513,-25.9805767,116.4895926,-1.0964358,-16.5175201,-56.3597400,-44.9448864,13.8954747,-12.9902884,-5.6437515,71.3703842,25.2180227,-41.2938002,-53.1391834,-32.5850426,8.9911895,12.9902884,31.9812582,1.0964358,-70.7893718,-33.8158440,-38.2031534,-15.5302365,-25.0526990,153.4053085,36.1823804,-34.2148630,-41.8672354,-19.1015767,22.8866643,0.9278777,20.8322796,-29.4955716,-43.4775137,-69.6645739,33.5126155,-45.4660092,26.3144585,-33.0350402,24.1541743,-42.6976421,0.0000000,-28.7642099,38.3752520,-7.0789372,-22.5438960,-20.2251989,34.3299964,19.4854325,4.3857431,-61.3507889,-33.8158440,-64.0464631,39.2342816,-28.7642099,183.7582306,-4.3857431,-22.4166344,-28.9850091,-57.3047302,25.3388069,-26.9084544,35.0859447,7.0789372,-33.8158440,-43.8212642,-1.6347617,5.5672664,-35.0859447,-40.1139773,-14.4925046,-12.3598438,21.2519025,-14.8460438,119.7709896,30.7002016,-22.4166344,-46.6980703,-43.8212642,5.7216661,-10.2066551,203.4466124,116.2221917,-83.7674233,-109.4989234,-38.2031534,78.4685632,-56.6005421,21.9287154,-63.7104346,-56.3597400,-4.4944886,25.3388069,-73.3023414,29.6037658,-31.8552173,-46.6980703,-79.7771734,21.2519025,-18.5575548,16.4465366,-27.1359259,-43.4775137,-41.5740199,-11.4433321,-23.1969435,27.4108943,-84.9472461,-53.1391834,-40.4503978,22.8866643,16.7017993)
推荐指数
解决办法
查看次数
识别和删除PCA和QQ图中的异常值
我有一个132 x 107的数据集,包括2个患者类型 - (患者1的33)和(患者2的99).
我正在寻找异常值,所以我在数据集上运行了pca,并使用以下命令完成了前4个组件的qqplots
pca = prcomp(data, scale. = TRUE)
plot(pca$x, pch = 20, col = c(rep("red", 33), rep("blue", 99)))
当我使用以下内容执行第二个组件的qqplot时:
qqPlot(pca$x[,2],pch = 20, col = c(rep("red", 33), rep("blue", 99)))
下图显示了2个明确的异常值 - 左下角的红点是患者1.
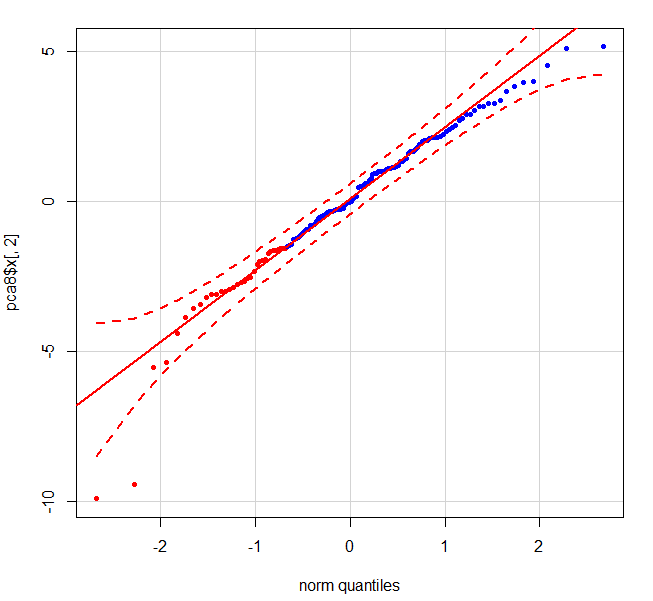
有没有直接的方法来计算数据中这些点的索引,以便可以删除它们?
推荐指数
解决办法
查看次数
x 轴上带有标签的绘图
我想要绘制 10 个不同分子的强度值的直线图,其中 x 轴为分子名称,y 轴为强度值。
我试过:
x = c("Mol 1","Mol 2","Mol 3","Mol 4","Mol 5","Mol 6","Mol 7","Mol 8","Mol 9","Mol 10")
intensity = c(428,409,388,378,373,140,137,138,139,144)
plot(x,intensity)
但是却返回了这个错误信息?
plot.window(...) 中的错误:需要有限的“xlim”值另外:警告消息:1:在 xy.coords(x,y,xlabel,ylabel,log) 中:强制引入的 NA 2:在 min( x) : min 没有非缺失参数;返回 Inf 3:在 max(x) 中:max 没有非缺失参数;返回-Inf
推荐指数
解决办法
查看次数
使igraph更加清晰易读
我有一个有69个顶点的有向图,如下所示。它是使用igraph软件包绘制的:
library(igraph)
ig <- graph.adjacency(data, mode="directed", weighted=TRUE)
plot(ig)
我正在寻求实现以下两点:
(a)将顶点隔开,并可能加长边缘,使其更易于阅读
(b)实际上,我的标签更长。是否可以使顶点变大而使文本变小以适应这个问题。
有任何想法吗?
这是我的数据:https : //www.dropbox.com/s/rtedrd1x1duqllj/data.Rdata?dl=0
推荐指数
解决办法
查看次数
在文本文件中读取的问题
我正在尝试使用以下代码将文本文件读入R:
d = read.table("test_data.txt")
它返回以下错误消息:
"Error in scan(file, what, nmax, sep, dec, quote, skip, nlines, na.strings, :
line 2 did not have 119 elements"
我试过这个:
read.table("man_cohort9_check.txt", header=T, sep="\t")
但它给出了这个错误:
"Error in scan(file, what, nmax, sep, dec, quote, skip, nlines, na.strings, :
line 43 did not have 116 elements"
我不明白出了什么问题?
推荐指数
解决办法
查看次数
